Loop mediated isothermal Amplification (LAMP) technique is one of the recent PCR techniques that is gaining importance for its application in molecular diagnostics. LAMP is a single tube PCR reaction carried out at constant temperature. LAMP is simple, affordable, cost effective, requires simple machinery and can be used in remote places by persons with little training.
LAMP PCR involves two main repetitive steps of elongation by loop primers:
- Self elongation of template from stem loop structure formed at 3’end and
- Elongation of new PCR product from the primers
The end product hence is not a single band but a series of concatamers of different sizes as DNA ladder or distinct band pattern on agarose gel. Bst DNA polymerase from Bacillus stearothermophilus helps in strand displacement and elongation of DNA.
Both RNA and DNA samples can be used as template for detection of the diseases. When RNA is the template either Bst enzyme with reverse transcriptase activity (Bst, Large fragment) or a separate reverse transcriptase is used along with normal Bst enzyme. This is called as RT - LAMP PCR. When DNA is the template it is called LAMP PCR.
For a successful LAMP reaction design of primers plays a major role. Primers are designed based on 6 regions in the target sequence. In LAMP assay 3 sets of primers are mainly used:
- Inner primers (Forward and Backward) —F1C+F2 and B1C+B2
- Outer primers (Forward and Backward) — F3 and B3
- Loop primers (Forward and Reverse) —FL and BL
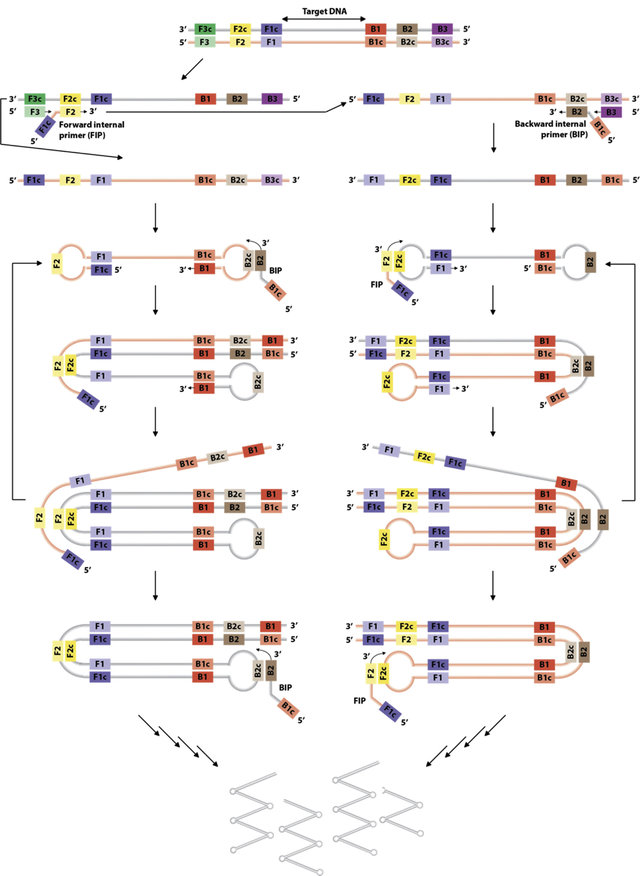
Figure: Loop-mediated isothermal amplification (LAMP) of DNA. The LAMP reaction employs several primers to target six distinct regions on the double-stranded DNA. The reaction is initiated upon binding of FIP to the F2c region of the DNA. DNA polymerase extends the DNA from the FIP. The primer F3 binds to its complementary region on the DNA to displace the newly synthesised DNA. Analogous reaction is performed by BIP and B3. Resulting displaced strands form 'dumbbell' structures. FIP and BIP bind to these dumbbell structures to synthesize a new strand that forms a loop due to the complementary of the regions. Elongation of the loop structure forms double-stem DNA loops that displaces the original strand, which can reform its dumbbell structure and undergo another round of amplification. BIP and FIP anneal to the double-stem DNA loops to prime further strand displacement. Repeated rounds of these steps amplify target DNA to produce DNA structures of various sizes. (Lubis et al. (2016) Trends in Food Science 7 Technology. 58:55)
Primer Melting Temperature (Tm), GC content, stability of primer ends, and secondary structure are the important factors to be considered. Tm of F1c & B1c, FL & BL should be 65 ℃ and Tm of F2 & B2, F3 & B3 should be 60 ℃. GC content should be between 50-60%, ends of primers should be stable with free energy of about -4 kcal/mol and 3’ends of primers should not be complimentary to avoid primer dimer formation. Primers should be designed to avoid secondary structures. Using primer explorer software one can design the lamp primers sets with less effort.
LAMP assay can be performed in less than one hour, including sample collection and subjecting to heat treatment or DNA extraction followed by adding the six primer mix and Bst DNA polymerase and incubating at constant temperature for 30 minutes. Some other parameters have to be optimized in addition to primers, such as enzyme concentration, reaction temperature, reaction pH, Mg2+ ion concentration, additives like Betaine, etc.,
LAMP PCR technique can be used for genetic testing, point of care diagnosis, detection of food and environmental samples in remote or resource poor areas. Positive reaction can easily be detected by naked eye either by turbidity or by non fluorescent dyes like phenyl red or fluorescent dyes like SYBR green or by an agarose gel in comparison to negative samples or control samples. Recently LAMP based kits for detection of diseases like Influenza, Tuberculosis, Malaria, etc have been developed. WHO recommends TB LAMP reaction as an alternative or follow-up test for microscopy in TB patients with signs and symptoms.
LAMP PCR can be used for detecting or diagnosing disease but can not be used for cloning purpose. Though there are few reports of multiplex LAMP PCR reaction, it is not well established to test large number of samples in short time. For avoiding non-specific amplification in negative samples several primer sets have to be synthesized and checked. Apart from these limitations, LAMP is a very promising technique to develop diagnostics for many diseases and can easily be used in remote areas.
References:
- Chia-Chen Chang, Chien-Cheng Chen, Shih-Chung Wei, Hui-Hsin Lu, Yang- Hung Liang and Chii-Wann Diagnostic Devices for Isothermal Nucleic Acid Amplification. Sensors 2012, 12(6), 8319-8337.
- World Health Organization .The use of loop-mediated isothermal amplification (TB-LAMP) for the diagnosis of pulmonary tuberculosis: policy guidance. 2016. ISBN: 9789241511186