 At a cellular level, there is always a homeostasis maintained between cell division and cell death. The process of growth and development involves an active balance of both of these mechanisms. Cell death occurs through three distinct mechanisms depending upon the molecular signal: Apoptosis, Necrosis, and Autophagy. All these processes have specific morphological and biochemical hallmarks that help them to get identified through different techniques. Also, these processes are crucial in a normal cellular state, while a heightened or reduced rate of these processes lead to diseased conditions. Autophagy involves intracellular protein degradation and is an important aspect of cell starvation, aging, and differentiation. Apoptosis is also known as programmed cell death, where a strict tightly controlled event making apoptotic bodies to promote cell death. Necrosis results into the release of cellular components in the extracellular space after cell destruction.
At a cellular level, there is always a homeostasis maintained between cell division and cell death. The process of growth and development involves an active balance of both of these mechanisms. Cell death occurs through three distinct mechanisms depending upon the molecular signal: Apoptosis, Necrosis, and Autophagy. All these processes have specific morphological and biochemical hallmarks that help them to get identified through different techniques. Also, these processes are crucial in a normal cellular state, while a heightened or reduced rate of these processes lead to diseased conditions. Autophagy involves intracellular protein degradation and is an important aspect of cell starvation, aging, and differentiation. Apoptosis is also known as programmed cell death, where a strict tightly controlled event making apoptotic bodies to promote cell death. Necrosis results into the release of cellular components in the extracellular space after cell destruction.
The characteristic morphological and biochemical features of apoptosis are:
- Chromatin condensation and cellular shrinkage
- Formation of apoptotic bodies (formation of membrane-bound cell fragments)
- DNA degradation by endonucleases
Running the samples on an agarose gel gives a crude idea of cell death by looking at the short or long length DNA fragments. However, it cannot provide a spatial information of the DNA fragmentation in a tissue or single cell population.
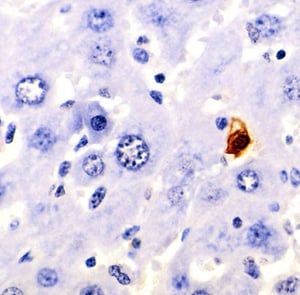
Terminal deoxynucleotidyl transferase (TdT) dUTP Nick- End Labeling (TUNEL) is the most common method used to assay the localization of DNA fragmentation caused by cell death. Apoptosis marked by nuclease activation and DNA fragmentation results into the formation of many DNA 3'-hydroxyl (OH) open ends, which facilitate end labeling of these fragments using fluorophores. The amount of fluorescence can be detected by fluorescence microscopy or flow cytometry. This method utilizes on the identification of exposed blunt ends of the double-stranded DNA in a template-independent manner. TdT enzyme used for this method allows the addition of labeled dUTPs at the 3'-OH terminal of the fragmented DNA. These labeled fragments can be identified in-situ using immunohistochemical techniques.
Incorporation efficiency of the modified dUTP decides the sensitivity of the TUNEL assay, which is overall influenced by the relative bulkiness of the labels.
There are many labels that can be attached by TdT to the DNA fragments such as
- Fluorescent-tagged dUTPs- FITC TUNEL assay
- biotinylated dUTPs
- DIG or digoxigenylated dUTPs
However, these tags are bulky and provide a lot of steric hindrances, therefore compromise with the sensitivity of the assay. Contrary to that, smaller labels such as bromine (BrdUTP) or an alkyne group (EdUTP) have higher sensitivity due to lesser steric hindrance. BrdUTP can be detected by using fluorophore or reporter enzyme-labeled antibodies and EdUTP detection is achieved by alkyne-azide click chemistry.
TUNEL assay has three simple processing stages:
- SAMPLE PREPARATION: TUNEL assay can be performed in any type of sample (tissue sections, cell suspensions, cell lines. It is essential to fix the sample with an appropriate fixative (1-4% paraformaldehyde) and permeabilized with either methanol or Tween-20. Samples can be stored at –20°C for further use or can proceed directly for the next step.
- TUNEL staining: The staining requires positive and negative controls along with the test cell samples. All the samples undergo incubation with TdT enzyme mixture for a specific time interval followed by labeling with the antibody mixture. DNA fluorochromes such as propidium iodide are used for background nucleus marking.
- ANALYSIS: The results can be observed through flow cytometry (cell suspension) or light or fluorescent microscopy ( adhered cells or tissue sections) depending upon the type of labeling.
TUNEL assay is very simple and handy method to perform, although it comes with few caveats which needs to take care of while performing experiments.
- As DNA fragmentation can result from various situations such as Necrosis, over-fixation, prolonged proteinase-K treatment, cell division, and tissue sectioning. Therefore, the baseline or threshold for every test and control samples needs to be set accordingly.
- Many cell types require overnight 70% (v/v) ethanol incubation at –20°C post-fixation for good incorporation efficiency.
- Apoptotic pathways differ from cell to cell, therefore TUNEL assay needs further confirmation by classic morphological assessments such as using DNA fluorochrome to confirm apoptotic bodies.
- The signal and the results of the TUNEL assay vary from sample to sample, therefore it requires optimization at two steps especially: Paraformaldehyde fixation and Proteinase K digestion. Over-fixation and over-digestion with proteinase K result into less signal in positive controls and test samples.