Carbohydrate affinity chromatography is a method of choice for purification of glycoproteins, lectins and other carbohydrate metabolite proteins. This affinity chromatography uses carbohydrate ligands such as carbohydrates, glycoproteins and carbohydrate matrices for purification. The matrix needs to be activated for covalent immobilization of the ligand. Various methods used for the coupling of ligands depending upon the activating reagents are as follows:
The divinyl sufone (DVS) coupling: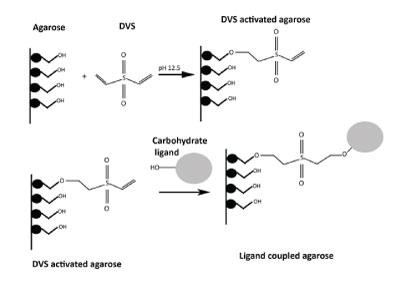
Diviny sulfone (DVS) is a bifunctional compound that is used to activate hydroxyl groups containing matrices. DVS activated matrices can covalently bind sugars and carbohydrates through the hydroxyl groups present on carbohydrates. The activation of matrix using divinyl suphone is rapid (70 min) at room temperature. This method not only introduces a highly reactive vinyl group but also a spacer arm to avoid steric hindrance of the bound ligands. The divinyl sulfone activated matrix is stable as suspension for 1 year at 4°C. The linkage chemistry of DVS coupled ligand is unstable in alkaline conditions, therefore acid elution of lectins is recommended when pH elution method is desired.
The epoxide coupling: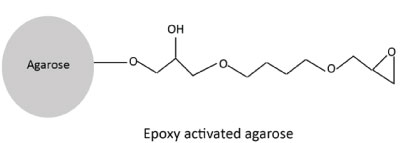
The epoxide method for coupling of ligand uses bifunctional compounds called oxiranes. Oxiranes such as 1,4-butanediol diglycidyl ether are immobilized on agarose matrix to form epoxide activated agarose. The epoxide activated agarose is unstable in aqueous solutions and so should be used immediately for coupling the ligand. The epoxide agarose form covalent bonds with the ligands that contain nucleophiles such as amino, thiol, hydroxyl and phenolic functional groups. Carbohydrates have free hydroxyl groups that interact with epoxides to form ether bonds. This coupling chemistry additionally provides a 12 atom hydrophilic spacer arm. Since the coupling conditions for carbohydrates require high pH and high temperature, this method is not recommended for silica and glass beads matrices.
The N-hydroxysuccinimide coupling:
In N-hydroxysuccinimide method, N-hydroxysuccinimide is used as coupling agent and this agent forms stable bonds with ligands containing amine groups. The stable amide bond is formed between the ligand and the coupling agent. It is used in coupling of glycoproteins or carbohydrate derivates that contain amino groups to the matrix. This coupling method in addition provides an eight carbon spacer arm.
The cyanogen bromide coupling:
This coupling method is used to immobilize ligands containing amino groups. This method is not suitable for coupling small molecules as it does not provide any spacer arm. This coupling method is suitable for immobilizing glycoproteins.
The thiopropyl coupling: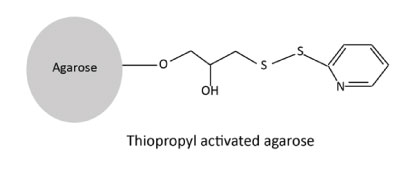
Thiopropyl activated matrix contains reactive 2-thiopyridyl disulphide groups that reacts reversibly with ligands containing thiol groups to form mixed disulfides. It is used to immobilize glycoproteins via free thiol groups present on glycoproteins. It can be used to immobilize iodoacetamidyl carbohydrate derivatives by alkylation of the thiol groups. The thiopropyl coupling method also provides a spacer arm.
The azlactone coupling:
Azlactone activated matrices are formed by co-polymerization of acrylamide with azlactone. This reacts with ligands containing amino groups and thus is suitable for binding of glycoproteins.